Two Centre for Heart Lung Innovation (HLI) researchers have been recognized with UBC’s 2025 Faculty of Medicine Distinguished Achievement Awards in the Excellence in Clinical or Applied Research category, which honours outstanding clinical or applied research and scholarly contributions.

Each year, the UBC Faculty of Medicine honours faculty members who demonstrate excellence in research, education and service, and who advance the Faculty’s mission of transforming health for everyone.
Dr. Pat Camp, an associate professor in the Department of Physical Therapy, leads the UBC Pulmonary Rehabilitation Research Laboratory. Her research focuses on developing patient-centered strategies to help people with chronic lung disease stay active and improve their overall health.
Dr. David Granville, a professor in the Department of Pathology and Laboratory Medicine, studies a group of enzymes called granzymes that play key roles in inflammation, tissue injury, and aging-related diseases. The Granville Lab is exploring how targeting these enzymes could lead to new treatments for conditions such as heart disease and chronic wounds.
Congratulations to Drs. Pat Camp and David Granville on this well-deserved award!
Read the original announcement by UBC here.
The first High School Student Science Week (HSSW) of 2025 ran from April 28 to May 2. Held twice a year, HSSW invites Grade 11 and 12 students from across the Lower Mainland to explore HLI’s labs and gain hands-on experience with molecular biology techniques. Experiences like these highlight how HLI connects students to real-world science and sparks curiosity across disciplines, continuing a tradition of mentorship and collaboration that stretches back to 1977.
As part of their visit, students had the unique opportunity to be exposed to biobanking and pathology. At the Bruce McManus Cardiovascular Biobank and James Hogg Lung Biobank, they handled preserved human heart and lung tissues, which was an eye-opening experience for many seeing real organs up close for the first time. In the histology session, they learned to prepare tissue samples by trimming paraffin blocks and using a microtome to make thin slices for microscopic analysis.
The group also toured several HLI research cores where staff introduced them to the tools and workflows behind scientific discovery. A highlight for many included a tour of the Anatomical Pathology Lab at St. Paul’s Hospital, where students examined other surgically explanted specimens.
The fall intake for HSSW will open in October. Stay tuned!


HLI’s Alumni Night on May 1 brought together former trainees and current students for an evening of career reflections, panel discussion and networking focused on paths in academia, industry, and more.
Former trainees at the Centre for Heart Lung Innovation (HLI) returned to St. Paul’s Hospital on Thursday, May 1, for the 2025 Alumni Night hosted by the Trainee Association at HLI (TAHLI) and the TAHLI Mentorship Committee. The event featured a panel discussion followed by a sushi dinner, bringing together 36 attendees — including current trainees, research staff, and five alumni who each made HLI part of their scientific career.
The panelists – Drs. Tillie Hackett, Dorota Stefanowicz, Kauna Usman, Kang Dong and Joshua Dubland – spoke candidly about their career journeys, the detours they encountered, and the soft skills that helped them succeed.
Dr. Dorota Stefanowicz shared her decision to move from academia to industry, driven by a need for better work-life balance. Dr. Tillie Hackett, now an HLI Principal Investigator and former HLI postdoc, highlighted the value of strong writing skills, not just for grant applications, but even in navigating everyday life, like supporting her child’s preschool journey.
Dr. Kauna Usman, who recently began a postdoctoral fellowship, described the excitement of stepping into a new research chapter: “It feels like having so many toys to play with,” she said.
Dr. Kang Dong, now a data scientist at Olink Proteomics, spoke about the importance of knowing when to move on from projects that no longer serve their purpose.
Dr. Joshua Dubland, a clinical assistant professor at UBC, reflected on his nontraditional journey, from studying music to working in industry to becoming a clinical scientist, and emphasized the power of persistence in the face of rejection.
When asked what soft skills are most important to them, the panelists offered various responses: writing and communication, adaptability, resilience, and knowing when to pause or pivot. Even with their different paths, they agreed that passion was the driving force behind their success, and that they are willing to work long hours when they are doing what they love with people they enjoy working with.
The evening left many attendees inspired and reassured that there is no single, correct roadmap to a fulfilling career in science.
As co-chair of TAHLI, Eric Xiang reflected:
“Navigating graduate training and job searching can be a challenging and non-linear journey, often filled with ups and downs. However, hearing the inspiring stories of our alumni fills me with optimism and motivation for the path ahead.”
“Navigating graduate training and job searching can be a challenging and non-linear journey, often filled with ups and downs. However, hearing the inspiring stories of our alumni fills me with optimism and motivation for the path ahead.”
— Eric Xiang, Co-Chair of TAHLI
Upcoming Events
The TAHLI Mentorship Committee is hosting an Early Career Night on Aug. 21 from 5–7 PM at the JHCC, one day before HLI Research Day. This event is a follow-up to Alumni Night, offering a second chance for those who missed it, and a new opportunity to connect with peers and returning alumni.
The 2025 Killam Teaching Prize recognizes HLI researcher Dr. Tillie Hackett for her commitment to inclusive teaching, mentorship and research leadership.
Dr. Tillie-Louise Hackett, a Faculty of Medicine Professor and lung health researcher at the University of British Columbia, has received the 2025 Killam Teaching Prize for her outstanding contributions to teaching, mentorship and research leadership.
The award is presented annually to faculty members who demonstrate exceptional dedication to teaching. Recipients are nominated by students, colleagues and alumni across the UBC Vancouver campus, and selected within each Faculty at UBC’s Vancouver campus.
“Receiving the Killam Teaching Prize is incredibly meaningful because it reflects the value of creating inclusive, engaging learning environments where students can thrive,” said Dr. Hackett. “It reinforces my belief that education is most powerful when it is collaborative, student-centred and grounded in real-world impact.”

A Tier 1 Canada Research Chair in asthma and COPD lung pathobiology and therapeutics, Dr. Hackett leads a research program focused on chronic respiratory disease. Her research focuses on improving the health of patients living with asthma, chronic obstructive pulmonary disease and idiopathic pulmonary fibrosis — people who face daily challenges with breathing. But her impact extends beyond the lab bench.
Over the past decade, Dr. Hackett has mentored dozens of trainees – six master’s and 10 doctoral students, eight postdoctoral fellows, six co-op students, 17 summer students, three clinical fellows and five BSc honours students. Her students have collectively won more than 150 awards, and many have gone on to pursue careers in medicine, academia and industry.
“What I find most meaningful is helping students connect scientific knowledge to practical, real-world applications,” she said.
“Whether it’s teaching inhaler techniques or guiding translational research projects, watching students gain confidence, ask thoughtful questions, and apply what they learn to improve patient care or push scientific boundaries is incredibly rewarding.”
— Dr. Tillie Hackett, HLI Principal Investigator
Dr. Hackett’s commitment to inclusive, student-centred teaching continues to shape the next generation of researchers.
“This award is a reflection of the vibrant, diverse community I have the privilege to teach and learn from,” she says.
“My students and trainees continuously inspire me with their dedication, curiosity and drive to make a difference. I’m proud to support them not only as learners but as future leaders in science and medicine.”
— Dr. Tillie Hackett, HLI Principal Investigator
Congratulations to Dr. Tillie Hackett on this well-deserved recognition!
Read More
New research from the Centre for Heart Lung Innovation (HLI) has identified inflammation in the lungs’ smallest airways as the cause of pulmonary long COVID-19.
A research team led by the HLI’s Director, Dr. Don Sin, used xenon MRI, an advanced imaging technology, and single-cell sequencing of lung samples, supported by the HLI’s biobank team, to uncover the underlying mechanism behind persistent respiratory symptoms in long COVID patients. These findings were published in three articles in the European Respiratory Journal.
What is long COVID?
Five years on from the beginning of the COVID-19 pandemic, many COVID patients continue to be impacted by residual symptoms lingering after the initial illness, a condition known as long COVID. Affecting approximately 10 per cent of people infected with COVID-19, long COVID is a multi-system disorder with diverse presentations, including brain fog, fatigue, joint pain, shortness of breath, wheeziness, and cough. These symptoms can significantly impact patients’ quality of life and are associated with increased health care costs.
The HLI research team focused on identifying the cause of pulmonary long COVID, which presents with persistent lung symptoms and accounts for approximately one-third of long COVID cases.
Detailed lung scans made possible with xenon MRI
Despite experiencing often-debilitating symptoms, 80 to 90 per cent of the long COVID participants tested by the HLI team showed normal results on standard lung function tests and CT scans.
“I distinctly remember a nurse who could no longer work after recovering from acute COVID. He had so much shortness of breath, even with just minimal exertion, that he had to go on long-term disability. And yet, his breathing test was normal. His CT scan was also normal,” says Dr. Sin.
The researchers suspected that these standard tests were not sensitive enough to detect the root cause of these symptoms, and so they turned to hyperpolarized xenon gas magnetic resonance imaging (xenon MRI), an advanced method that enables three-dimensional imaging of lung function.
In 2019, with support from St. Paul’s Foundation and the Canada Foundation for Innovation, St. Paul’s Hospital obtained a specialized MRI and hyperpolarizer. Together, these devices enabled the use of inhaled hyperpolarized xenon gas to image the lungs’ airways and track the flow of oxygen into the bloodstream, a process known as gas exchange. Hyperpolarization enhances the xenon’s magnetism, making it more visible to the MRI.
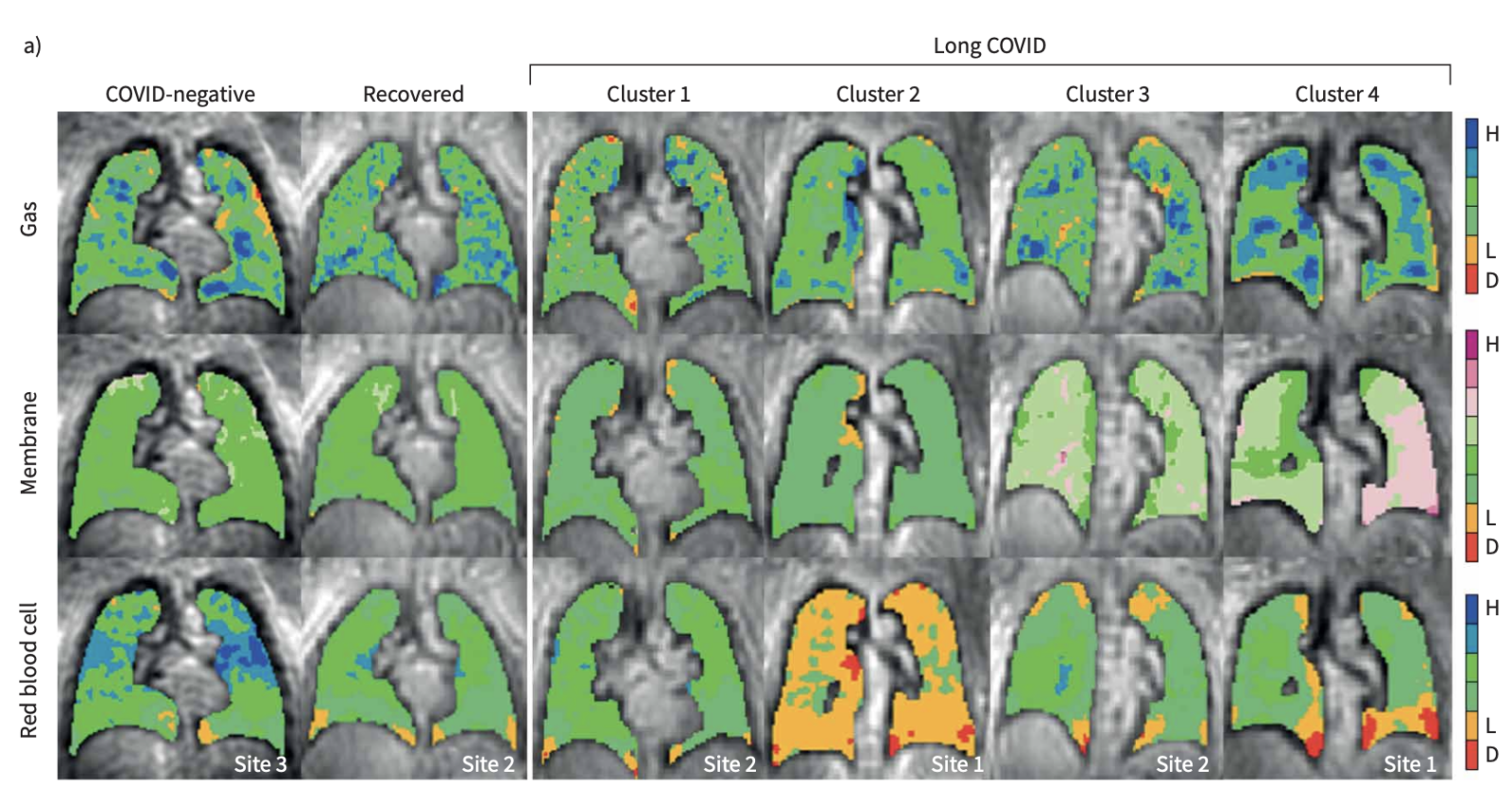
“It’s particularly the gas exchange component that makes xenon MRI so unique,” says Dr. Rachel Eddy, Director of the HLI’s MRI core who launched the centre’s xenon MRI research program. As xenon follows the same gas-transfer pathways as oxygen, xenon MRI is able to image the separate components of gas exchange, allowing researchers to identify where in the lungs a problem is occurring.
By analyzing combined data from the HLI, Duke University, and the University of Kansas Medical Centre from xenon MRI scans of a large cohort of long COVID patients, the researchers identified four clusters, or subgroups, of pulmonary long COVID, which were characterized by different types of gas exchange abnormalities.
The researchers determined that pulmonary long COVID is primarily driven by gas exchange problems in the small airways, where oxygen enters the bloodstream, which cannot be visualized in traditional scans or lung function tests.
“That’s what hyperpolarized xenon gas MRI does. It lets us see beneath the surface,” says Dr. Sin.
Single-cell profiling identified inflammation in small airways
To determine the cellular and molecular drivers of these abnormalities, a subset of patients who had participated in the xenon MRI study underwent bronchoscopies to take tissue samples from their small airways, which were analyzed with single-cell RNA sequencing.
Because single-cell sequencing requires fresh, living tissue, the research team faced logistical challenges in transporting the samples from St. Paul’s Hospital to the University of British Columbia, where the sequencing equipment is located. To ensure the cells survived the journey, they utilized the expertise of the HLI biobanking team to process and prepare the samples.
“The biobanking facility is much more than just keeping things in the freezer. It’s actually developing protocols and methods to enable the use of human volunteer tissues for research,” says Dr. Sin. The HLI is home to several influential biobanks, including the James Hogg Lung Biobank and the Bruce McManus Cardiovascular Biobank.
The single-cell sequencing found that the pulmonary long COVID patients had neutrophilic inflammation in the small airways. Neutrophils, immune cells that normally help fight infection, were continuing to trigger an immune response, even though the virus was gone.
“They’re like dirty bombs. They come in to kill the bacteria or viruses,” says Dr. Sin. “Once all the pathogens are killed off, then the body should shut down the recruitment of these cells into the small airways.” In pulmonary long COVID patients, these cells remain, causing damage to the small airways and driving the symptoms of pulmonary long COVID.
Inflammation is likely to resolve
A few key questions remain, such as why some people develop long COVID while others recover fully. It is still unknown why these inflammatory cells persist, or if this phenomenon could occur with other viruses like influenza or RSV.
“We don’t think this is specific to coronavirus. If a virus gets in deep enough into your airways, we think that this can trigger, in some individuals, a persistent response for a period of time,” says Dr. Sin.
Encouragingly for pulmonary long COVID patients, Dr. Sin and his team believe that this inflammation, which is relatively mild, will likely resolve on its own over time, provided it isn’t exacerbated by factors like smoking, exposure to wildfire smoke, or additional COVID-19 infections.
“Prevention, breathing in clean air, refraining from smoking and dusty environments, those are, I think, very important preventative measures. If patients keep on doing that, over time we think this inflammation will settle on its own,”
— Dr. Don Sin, HLI Director
Read more
- Cluster analysis to identify long COVID phenotypes using 129Xe magnetic resonance imaging: a multicentre evaluation
- Single-cell sequencing reveals cellular landscape alterations in the airway mucosa of patients with pulmonary long COVID
- A proposed approach to pulmonary long COVID: a viewpoint
This article was originally written by Grace Jenkins and published on March 24, 2025 through Providence Research. Read the article on Providence Research here.
On this year’s National Indigenous Languages Day (March 31, 2025), we highlight Dr. Pat Camp’s important work in incorporating Indigenous language into culturally safe healthcare and research practices.
Dr. Pat Camp, a Principal Investigator at the Centre for Heart Lung Innovation (HLI), reflects on her research helping people manage chronic lung conditions through exercise, education, and support – a process called pulmonary rehabilitation.
Through collaboration with Carrier Sekani Family Services, Dr. Camp shares how her team is learning to respectfully incorporate Indigenous knowledge into pulmonary rehabilitation research. This includes using the Carrier (Dakelh) language to name body parts—an approach to make pulmonary rehab more culturally safe, respectful, and healing.
“There’s a lot more that we can do”
— Dr. Pat Camp, HLI Principal Investigator
Watch Dr. Pat Camp’s Story
Learn More
Learn about National Indigenous Language Day:
Other links:
New research on low-carb, high-fat diets earns a spot among the five most-read articles in one of cardiology’s leading journals.
A study led by Dr. Iulia Iatan and Dr. Liam Brunham at the Centre for Heart Lung Innovation (HLI) has been recognized as one of JACC: Advances’ Top 5 Most-Read Articles of 2024. Published in one of the most prominent cardiovascular journals, the study’s ranking underscores its impact and the global interest in its findings.
“It is an honour to see our study recognized among the most-read articles in JACC: Advances for 2024. This reflects the growing interest in understanding how dietary choices impact long-term heart health,” said lead author Dr. Iulia Iatan, a clinician-scientist and former postdoctoral fellow at UBC’s Centre for Heart Lung Innovation and the Healthy Heart Program Prevention Clinic at St. Paul’s Hospital.
“To our knowledge, this was one of the first studies to show an association between low-carbohydrate high-fat diets, increased cholesterol, and higher risk of cardiovascular events.”
— Dr. Iulia Iatan, HLI’s former postdoctoral fellow
“I hope these findings encourage both the public and healthcare professionals to consider the long-term effects of these diets, and also inform clinical practice and public awareness on the importance of heart-healthy nutrition.”
Trendy “keto-like” diet under scrutiny
Low-carbohydrate, high-fat (LCHF) diets have gained popularity for various health reasons, including weight loss and diabetes management, but their long-term effects on heart health have been uncertain. This study found that “keto-like” diets are linked to higher cholesterol levels and a twofold increase in major heart problems, including heart attacks and strokes over the next decade. The risk was highest for those with a genetic predisposition to high cholesterol.
To investigate this link, Dr. Iatan and the team analyzed data from the UK Biobank, tracking 305 participants who followed an LCHF diet—defined as <25% of daily calories from carbohydrates and >45% from fat—over an average of 12 years. They examined lipid levels and cardiovascular events, comparing outcomes to those on normal, standard diets.
A further finding from the study was that the risk of severely elevated cholesterol was highest among those individuals with an elevated polygenic risk score, suggesting a genetic predisposition to diet-induced increases in cholesterol in some people.
These findings emphasize the need for caution among followers of this dietary pattern, especially for those with high cholesterol or a personal or family history of heart disease.
What’s next?
Dr. Brunham and colleagues plan to continue examining how genetic differences influence response to various dietary patterns and their effects on heart health. Future research will also explore whether genetic testing can help create more personalized dietary recommendations and improve understanding of variations between individuals in response to different diets.
A big moment for HLI
This recognition reinforces HLI’s leadership in cardiovascular research and its role in providing strong, evidence-based guidance on health and disease prevention. With this study ranking among JACC: Advances’ most-read of the year, HLI continues to drive research that informs medical practice and public health.
“This is a great recognition of the quality of cardiovascular research taking place at HLI and our centre’s leadership in the field of lipid disorders.”
— Dr. Liam Brunham, HLI’s principal investigator
Further Reading
- Read the full study: JACC: Advances
- Check out Dr. Iulia Iatan’s CNN feature: CNN News Video
- Watch the 3-minute summary by Dr. Liam Brunham: PACE-CME Video
Media Coverage
Read our previous article on this study: ‘Keto-Like’ Diet May be Linked to Increased Risk of Heart Disease
Congratulations to Dr. Honglin Luo, Principal Investigator at the Centre for Heart Lung Innovation (HLI) and Professor in UBC’s Department of Pathology and Laboratory Medicine, on her nomination for the YWCA Women of Distinction Awards in the Research, Sciences, and Technology category.

Dr. Luo is an internationally recognized leader in viral pathogenesis and virotherapy, a dedicated mentor and role model to women and minority groups, and a highly-respected member of the scientific community. She has published over 125 peer-reviewed articles in top-tier journals and has been recognized by Expertscape as Canada’s leading experts in enterovirus research, ranking in the top 0.21% of 25,997 experts worldwide.
Her innovative research focuses on developing therapies for viral myocarditis, neurodegenerative diseases such as ALS, and cancer. Her lab’s pioneering work in oncolytic virotherapy—harnessing engineered or naturally occurring viruses to target and destroy cancer cells—has led to two successful patents for lung and breast cancer treatments.
Over the course of her career, Dr. Luo has demonstrated a sustained commitment to mentorship and service, dedicating herself to creating an equitable, diverse and inclusive environment. She actively supports the academic and research needs of her trainees—85% of whom are visible minorities—and encourages social dialogue and gender equity.
“Seeing Change Inspires Change”—YWCA
The YWCA Women of Distinction Awards celebrate women whose contributions drive innovation and mentorship. Dr. Luo’s nomination highlights her leadership in scientific research and her impact as a mentor, inspiring the next generation of women in STEM.
“As a woman in science, I have experienced the challenges of gender disparity in research. I advocate for Gender Equity because promoting equal opportunities in science empowers more women to lead and shape the future of technology.”
—Dr. Honglin Luo, HLI principal investigator
Show Your Support—Vote for Dr. Luo!
The YWCA Women of Distinction Awards also include a Connecting the Community category, where the public can vote for their favourite nominee. Cast your vote for Dr. Luo today before April 1, 12:00 PM!
🔗 Vote now: ywcavan.org/CCA-vote
